Vol. XXXIV Issue 1
Article 4
DOI: 10.35407/bag.2023.34.01.04
ARTÍCULO ORIGINAL
The importance of deep genotyping in crop breeding
La importancia de la genotipificación profunda en el
mejoramiento genético
Zambelli
A.1,2
1 Facultad de Ciencias Agrarias, Universidad
Nacional de Mar del Plata, Ruta 226 km 73.5, (7620) Balcarce, Buenos Aires,
Argentina
2 CONICET
* Corresponding author: Andres Zambelli
andres.zambelli@mdp.edu.ar ORCID 0000-0003-2057-4653
ABSTRACT
One of the greatest challenges
facing humanity is the development of sustainable strategies to ensure food
availability in response to population growth and climate change. One approach
that can contribute to increase food security is to close yield gaps and enhancing
genetic gain; to such end, what is known as “molecular breeding” plays a
fundamental role. Since a crop breeding program is mainly based on the quality
of the germplasm, its detailed genetic characterization is mandatory to ensure
the efficient use of genetic resources and accelerating development of superior
varieties. Deep genotyping is an essential tool for a comprehensive
characterization of the germplasm of interest and, fortunately, the technology
is now accessible at a reasonable cost. What must be ensured is the correct interpretation
of the genotypic information and on that basis develop efficient practical molecular
crop breeding strategies that respond to the real needs of the breeding
program.
Key words: Breeding population, Genetic resources,
Marker assisted selection, Single Nucleotide Polymorphism (SNP)
RESUMEN
Uno
de los mayores desafíos que enfrenta la humanidad es el desarrollo de
estrategias sostenibles para asegurar la disponibilidad de alimentos en
respuesta al crecimiento de la población y el cambio climático. Un enfoque que
puede contribuir a aumentar la seguridad alimentaria es cerrar las brechas de
rendimiento y mejorar la ganancia genética; para tal fin, lo que se conoce como
“mejoramiento molecular” juega un papel fundamental. Dado que un programa de
mejoramiento de cultivos se basa principalmente en la calidad del germoplasma, su
caracterización genética detallada es fundamental para garantizar el uso
eficiente de los recursos genéticos y acelerar el desarrollo de variedades
superiores. La genotipificación profunda es una herramienta esencial para una
caracterización integral del germoplasma de interés y, afortunadamente, en la
actualidad se puede acceder a la tecnología a un costo razonable. Lo que debe
asegurarse es la interpretación correcta de la información genotípica y sobre
esa base desarrollar estrategias eficientes y prácticas de mejoramiento
molecular de cultivos que respondan a las necesidades reales del programa de
mejoramiento.
Palabras clave: Población de
mejoramiento, Recursos genéticos, Selección asistida por marcadores, Polimorfismo
de nucleótido único (SNP)
General Editor: Elsa Camadro
Received: 04/25/2022
Revised version received: 07/11/2022
Accepted: 07/14/2022
INTRODUCTION
Crop breeding is a long-term
process that usually takes around ten years to develop and release a new
variety. Crop breeding is a large-scale logistical operation involving
thousands to hundreds of thousands of plants in the initial line fixation
stage, but numbers are greatly reduced to a small number of selected advanced breeding
lines by the end of the process: approximately 99% of the original starting
material in a breeding program is rejected and discarded (Lenaerts et al.,
2019). The Food and Agriculture Organization of the United Nations defined
modern plant breeding as “the act of using genetic diversity to improve the
agronomic performance of plants conducted as a formal endeavor and according to
scientific principles” (FAO, 1997). Cooper et al. (2014) defined modern
plant breeding as an integration of quantitative genetics, statistics, gene-to-phenotype
knowledge, and development models, applied to understand the functional
diversity of germplasm (Smith et al., 2015).
Crop improvement in a context of
continuous population growth and with climatic changes affecting agronomic
production has become a major global concern (Hickey et al., 2019).
Faced with these threats, current crop improvement strategies are unlikely to achieve
genetic gains that satisfy the demand for food both in terms of quantity and
quality. In addition, radical changes derived from climate change are causing heat
stress and drought, which leads to significant yield losses, so plant breeding
strategies need to be adapted to increase their efficiency.
The application of molecular
genetics in crop improvement has spread significantly since the appropriate use
of the so-known “molecular breeding” (i.e., genotype-based approaches) has
demonstrated to contribute to increase genetic gain with a highly favorable
cost-benefit ratio (Ismail and Horie, 2017; Xu et al., 2017;
Bailey-Serres et al., 2019). The correct choice of genotyping technology
allows a fine genetic characterization of germplasm, assisted selection, as
well as the implementation of genomic selection strategies.
Lack of in-depth analysis when
implementing a molecular breeding strategy can lead to failure, generating many
undesirable results and discouraging breeders from using the technology.
Consequently, before implementing a molecular breeding strategy, a serious
analysis of its advantages and disadvantages is strongly recommended, taking
into consideration the DNA technology of choice, the genetic diversity of the germplasm,
the architecture of the traits of agronomic interest to be improved, and the
resources demanded (Zambelli, 2019; Bohar et al., 2020).
GENETIC CHARACTERIZATION OF GERMPLASM
The configuration of an efficient
molecular breeding strategy must begin with a comprehensive genetic characterization
of the germplasm of interest through deep genotyping. Once characterization is
complete, the next challenge is to identify useful applications of genotype-based
technology to increase genetic gain. Genetic characterization is relevant for
germplasm management and agronomic use of both agricultural crops and their
respective wild relative species. The use of genomic tools is today technical
and practically more accessible than before, mainly due to the development of
next-generation sequencing (NGS) technologies and the reduction of their
application costs (Wu et al., 2014; Dempewolf et al., 2017;
Milner et al., 2019; Sansaloni et al., 2020; Fu et al.,
2021).
The topics to be listed when
addressing the genetic characterization of germplasm include SNP deep genotyping,
genetic diversity, genetic relationships, linkage disequilibrium, association
mapping, and population structure.
SNP deep genotyping
Different types of molecular
marker systems have been used for genotyping applied in plant breeding:
restriction fragment length polymorphisms (RFLPs), randomamplified polymorphic
DNAs (RAPDs), amplified fragment length polymorphisms (AFLPs), Diversity Arrays
Technology (DArT) and simple sequence repeats (SSRs). However, currently the
most advanced and commonly used marker systems are single nucleotide polymorphisms
(SNPs). Their abundance in genomes and the achievability to adapt them to
automated platforms have expanded access to deep genotyping at reasonable
costs, making SNPs the most widely adopted marker system for different genomic
applications (Mondini et al., 2009). With the increasing throughput of
NGS technologies, de novo and reference-based SNP discovery are today
feasible for most crop species.
When applying NGS two variables
need to be attended: coverage and sequencing depth. Coverage indicates the average
number of reads that cover a specific target genomic region, describing a
relationship between the number of reads and a reference region, and can be expressed
in terms of average coverage (for example, 10X means that on average the target
regions are covered by 10 reads). Instead, sequencing depth describes the absolute
number of total usable reads produced by sequencing, usually expressed in number
of reads (in millions). Depending on the experimental objective of interest,
coverage can vary from the entire genome, one locus, or random nucleotide
positions.
There are several genotyping
methods available which are generally offered by commercial parties for which
only tissue samples need to be sent for DNA extraction. Widely adopted
genotyping options fall into three categories: whole genome resequencing (WGR),
reduced representation sequencing (RRS), and SNP arrays (Scheben et al.,
2017). WGR and RRS methods are based on NGS technologies and bioinformatics
pipelines that align reads to a reference genome and call both SNPs and
genotypes (Scheben et al., 2017; Pavan et al., 2020). WGR differs
from RRS in the absence of a stage of reduction of genome complexity. RRS
usually employs restriction enzymes (RE) to digest genomic DNA prior to sequencing
(method identified as RE-RRS) giving rise to genotyping-by-sequencing
technology or GBS (Elshire et al., 2011).
SNP arrays rely on
allele-specific oligonucleotide (ASO) probes (including target SNP loci plus
their flanking regions) fixed on a solid support, which are used to interrogate
complementary fragments from DNA samples and infer genotypes based on the interpretation
of the hybridization signal. The two leader manufacturers (Affymetrix™ and
Illumina™) had developed 46 SNP arrays for 25 crop species with several markers
ranging from 3K to 820K, although for their routine application in the
molecular breeding of the most prominent field crops, arrays of 25-50K are usually
chosen (Rasheed et al., 2017).
The use of WGR, at least for the
moment, is not considered financially feasible in large genome crops such as
corn (2.5 Gbp), barley (5 Gbp), and wheat (17 Gbp). However, the final decision
on the convenience of making the investment will depend on the commercial importance
of the crop, the added value of the trait of interest and the expected net
return. For most of molecular breeding applications, deep genotyping by using
GBS or SNP arrays is recommended as they allow a satisfactory balance among the
number of SNP loci genotyped, quality data, and costs.
In GBS technology, the
allele-calling does not require a reference genome, offering an unbiased method
to assess genetic diversity in a large collection of accessions, especially in
orphan crops because SNP discovery and genotyping can be done simultaneously
with less bias toward genetic backgrounds (Rasheed et al., 2017; Darrier
et al., 2019). When the germplasm includes commercial materials combined
with exotic materials, GBS would be the most appropriate genotyping. The
disadvantage of using SNP arrays is the risk that some SNPs may not be
informative for all individuals. Since the ASO probes immobilized in the array
are fixed and predefined (identified from a restricted set of genotypes, mostly
public) the proportion of useful SNP for capturing the genetic diversity of the
germplasm of interest cannot be predicted. Therefore, one of the limitations to
work with SNP arrays is the ascertainment bias since they cannot identify
marker-trait associations for SNPs that were not present in the population used
for array development (Frascaroli et al., 2013; Lachance and Tishkoff,
2013; Rasheed et al., 2017; Negro et al., 2019). Contrarily, GBS
ensure that all SNPs discovered will be informative for all the sequenced
genotypes of interest, producing high-quality polymorphism data. Although actual
relative costs vary with the number of samples and the SNP density required, is
widely considered that pricing of genotyping by GBS is lower than SNP arrays (Li
et al., 2015; Pavan et al., 2020). An extra complexity of GBS
respect to arrays is the necessity of library preparation and bioinformatics
analysis (Elshire et al., 2011; Li et al., 2015; Sansaloni et
al., 2020; Fu et al., 2021). The good news is that there are many
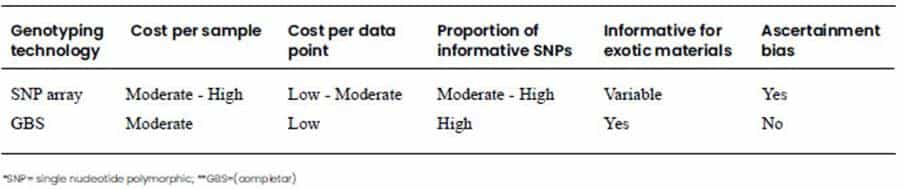
companies that provide the service at reasonable prices. In Table 1, the main
features of both genotyping technologies are summarized.
Tabla 1. Comparison of the main features
of genotyping by SNP* array and GBS** technologies)
Once genotype information is
collected (independently of the technology used), an adequate filtering
criterion considering different indicators should be followed to define the
high-quality genotyping dataset to avoid inaccuracies and bias in downstream
analyses. The presence of SNP loci with a high rate of missing data is
considered a feature of inaccurate genotype calls, so those SNPs should be
excluded from the analysis. SNP loci characterized by excessive heterozygosity
should also be excluded, as they are indicative of technical artifacts or
paralogous/repetitive regions that could not be distinguished through the
genotyping procedure. SNP loci displaying very low frequency alleles may derive
into genotyping errors and provide poor statistical power to reveal association
with phenotypic traits or establishing relative kinship. Thus, the recommended conditions
that SNPs should meet are: (i) up to 10% of missing genotype calls; (ii) up to
10% of heterozygous calls (assuming inbred lines are being genotyped); (iii) the
number of heterozygous calls does not exceed the number homozygous minor allele
counts; and (iv) minor allele frequency (MAF) > 0.05 (Wu et al.,
2014; Darrier et al., 2019; Milner et al., 2019; Pavan et al.,
2020).
Genetic diversity and genetic relationships
Productivity of most of field
crops remains far below the potential due to several factors such as access to high
quality seeds, irrigation, and fertilizers, abiotic stresses, high incidence of
pests and diseases, and weeds. However, genetic improvement provides an approach
to address some of these constraints, but largely depends on the availability
of genetic diversity, systematic classification, and efficient use of the available
germplasm.
A high-impact activity that
contributes to improving germplasm management and utilization is the analysis of
patterns of genetic diversity and population structure, which is important for
broadening the genetic basis and therefore, to establish successful commercial
breeding. Breeders demand a detailed genetics information of germplasm in order
to (i) define core subsets of germplasm for specific traits, (ii) select
parental combinations for developing progenies with maximum genetic variability
for further selection, (iii) identify genetic duplicates for better germplasm
management, (iv) enhance the search for unique germplasm with traits of
breeding targets for better varietal development, (v) describe heterotic groups
(Mohammadi and Presanna 2003; Reif et al., 2003; Flint-Garcia et al.,
2009; Ertiro et al., 2017; Ellis et al., 2018; Jeong et al.,
2019; Singh et al., 2019; Sansaloni et al., 2020).
The assessment of genetic
diversity within and between plant populations can be performed by using morphological
features, biochemical characterization of allozymes, and DNA markers. DNA
markers offer several advantages over phenotype-based alternatives as they are
stable and detectable in all tissues regardless of growth, differentiation, or
development stage and additionally, are not confounded by environmental, pleiotropic,
and epistatic effects. The availability of low cost and high throughput SNP
platforms facilitate genetic characterization of germplasm contributing to study
the amount and distribution of genetic variation they contain, arising as a
potent tool both for hybrid breeding and inbred breeding. Use of genotype data to
study genetic diversity can be mainly performed by calculation of population
genetics parameters and analysis of genetic relationships among samples (Govindaraj
et al., 2015).
The measuring of genetic
diversity is based on comparisons of individual genotypes within and between populations.
The analysis starts with the construction of a genotype matrix, sample × sample
pairwise and the calculation of the genetic distance (or similarities) that can
be done by different statistical methods, such as: (i) Nei and Li’s
coefficient, (ii) Jaccard’s coefficient, (iii) simple matching coefficient, and
(iv) modified Rogers’ distance (Mohammadi and Presanna, 2003).
The two main ways of analyzing
the resulting matrix are principal coordinate analysis (PCoA) and dendrogram (or
clustering tree diagram). PCoA is used to produce a 2- or 3-dimensional scatter
plot of the samples such that the distances among the samples reflect the
genetic distances among them with a minimum of distortion. The second approach
is to produce a dendrogram where samples are grouped in clusters according to
their genetic similarity. Different algorithms were used for clustering,
including Unweighted Pair Group Method with Arithmetic Averages (UPGMA),
neighbor-joining, and Ward’s method (Govindaraj et al., 2015).
Cluster analysis is of great help
for breeders in defining which genotypes should be crossed to develop breeding populations
that increase the chances of obtaining novel allelic combinations and to
reverse or mitigate the genetic erosion. Besides, the analysis of genetic relationship
is particularly useful when identifying the best materials to quickly integrate
them into an eroded germplasm pool through exchange, purchase, or inlicensing germplasm
(Beckett et al., 2017; Leitão et al., 2017; Vendelbo et al.,
2020). Different genetic materials, such as elite lines, ecotypes, landraces,
subspecies, or wild relatives, are potential useful sources of genetic variation.
Lack of genetic variation for traits of interest within the domesticated
genetic pool, imposed a greater exploration of crop wild relatives (CWR). Thus,
breeders in barley, maize, wheat, rice, sorghum, and soybean (among other
species) reported a lack of variation for traits of interest within the
domesticated germplasm, being exploration of CWR a feasible approach to mitigate
the genetic erosion (Pourkheirandish et al., 2020). Dempewolf et al. (2017)
reviewed how CWR contributed to the development of improved crop varieties by
crossing them with wild species carrying beneficial allelic variation for
traits. Private industry has valued the diversity of CWRs and landraces, which sometimes
is preferred as an alternative to the use of transgenic technology associated
with high regulatory costs and often resisted by consumers (Dempewolf et al.,
2017). The proper use of GBS constitutes a powerful tool to reveal and measure
the genetic variation contributed by wild species, a previous step required for
its potential use in crop improvement (Xu et al., 2017).
Existence of heterotic parental
gene pools constitutes the cornerstone in hybrid breeding programs as the prerequisite
for achieving a high heterosis effect in hybrid crosses. Hybrid crop breeders
evaluate the germplasm to assign inbred lines into distinct heterotic groups by
studying combining ability, mainly based on grain yield. However, the use of
molecular markers for genetic characterization of inbred lines can complement
and fine-tune the combining ability data. Genetic distance estimates contribute
to the assigning of genotypes to heterotic groups and the exploitation of
complementary lines which maximize the outcome of hybrid breeding programs (Wu et
al., 2014; Xu et al., 2014; Zhao et al., 2015; Beckett et
al., 2017; Labroo et al., 2021; Silva et al., 2021).
Thus, plant breeding community
has recognized that exploitation of genetic variability by conventional plant breeding
in combination with genomics approaches have contributed to developing high
yielding varieties or hybrids reducing the breeding cycle (Varshney et al.,
2005, 2021).
Linkage disequilibrium
Selection during crop breeding
has caused a dramatic loss of genetic diversity in many genome regions of modern
varieties. For instance, in major cereals and sunflower, reductions in
diversity of 30-40% and 40- 50%, respectively were estimated (Buckler et al.,
2001; Whitt et al., 2002; Liu and Burke, 2006). Thus, it can be assumed
that CWR for most crop species may have retained genetic information before
domestication and artificial selection. Linkage disequilibrium (LD) refers to the
non-random association of alleles at different loci (SNPs). LD is a common
variable in population genetics and evolutionary biology, used among others, to
map quantitative trait loci, estimate effective population size and past
founder events, or to detect genomic regions under selection (Lucek and Willi,
2021).
Both D′ and r2 statistics
have been widely used to quantify LD, differing in how they are affected by marginal
allele frequencies and small sample sizes. To identify SNPs significantly
associated with phenotypic trait variation, r2 is the most relevant LD
measurement. In small populations, the effects of genetic drift result in the
consistent loss of rare allelic combinations, which increase LD levels. When
genetic drift and recombination are at equilibrium, r2=1/(1+4Nec),
where Ne is the effective population size and c is the
recombination fraction between sites (Flint-Garcia et al., 2003). Ne is
one of the most important indicators in population genetics for describing the
magnitude of genetic drift, inbreeding, and assessing genetic diversity. The
smaller the effective population size, the faster the population will become inbred
and thus no longer respond to selection (Cobb et al., 2019). Ne is
an important parameter that helps to quantify the magnitude of genetic drift
and inbreeding. Thus, it is highly recommended that breeders actively calculate
and monitor Ne through successive breeding cycles to ensure the
long-term viability of their breeding programs. Knowledge of Ne helps
both, to design efficient selection and, if necessary, to modify parental
combinations that maintain or increase genetic variation to ensure the
identification of future superior candidates. In larger populations more
recombination events occur for which it is expected to have lower levels of LD.
Ne can be estimated by using both pedigree and marker data, however the
latter is presently preferred (Wang, 2016).
In practice, Ne is
directly related to the effective number of loci (Me), which can be
defined as the number of independent loci that gives the same variance of
realized relationship as obtained in the more realistic situation calculated by
Me=(2NeL)/log(4NeL), where L is the genome size in Morgan.
A larger Me (due to a larger Ne, L, or both) will require a
proportionally larger number of markers to capture the relatedness structure of
the population (Goddard, 2009; Wang, 2016). If a true functional polymorphism
contributes a fraction of the total trait variation, h2 q, and has a LD value
of r2 with another SNP, then the trait variation that can be explained
by this SNP will be r2 × h2 q. A similar inference cannot be made
using D′ (Zhu et al., 2008). Typically, r2 values of
0.1 or 0.2 are used to describe the LD decay. For instance, in soybean, a mild
decline in LD over distances as great as 50 kbp was described (Zhu et al.,
2003), whereas in rice it was found that LD approaches r2 = 0.10 for
distances from around 100 kbp (Garris et al., 2003). In contrast, in
maize and cultivated sunflower r2 declines to <0.10 within around 1
kb (Remington et al., 2001; Liu and Burke, 2006).
Measuring of the pattern and
extent of LD are influenced by different factors such as mating type, genetic
drift, gene flow, selection, mutation, population substructure and relatedness,
and ascertainment bias (Flint-Garcia et al., 2003). For instance,
domestication can induce population bottlenecks producing higher levels of LD
(slow decay). Similarly, the increase in homozygosity associated with
self-fertilization reduces the effective recombination rate, resulting in
elevated LD (rapid decay) across the genome (Nordborg, 2000) or localized
around the targeted loci (Clark et al., 2004). Conversely, gene flow and
recombination are predicted to reduce LD (Slatkin et al., 2008).
Association mapping and population structure
Investigating the magnitude of LD
decay determines the resolution of association mapping (AM) and marker-assisted
breeding for which studying the LD pattern contribute to estimate the required
numbers of SNPs. AM (also known as LD mapping) is a method of mapping
quantitative trait loci (QTLs) using historical meiotic recombination events
performed over several generations to associate phenotypes with genotypes in
large germplasm populations. AM provides relevant information into the genetic
basis of complex traits and is a valued approach to identify the genes
underlying agronomically important traits. AM is based on the LD between
molecular markers (SNPs) and functional loci, requiring detailed understanding
of the pattern of LD. AM of a trait-associated allele is based on the slow decay
of LD with closely linked markers (Slatkin et al., 2008; Zhu et al.,
2008). For instance, resequencing of cultivated and wild soybeans showed that
LD decayed relatively slowly; given the high LD, only a small subset of SNPs
would be required for marker-assisted breeding. However, the high LD introduces
limitations for association studies using genetic populations (Lam et al.,
2010).
Germplasm with a recombination
history producing a limited gene flow can result in a structured breeding population
with an uneven distribution of alleles across subgroups. Therefore, the use of
AM in such stratified populations may lead to non-functional and spurious associations.
However, statistical analysis that estimate the effects of population
structure–induced linkage disequilibria allowed to expand the proper use of AM (Pritchard
et al., 2000).
The domestication of crops has
generated new population structures, some of which were geographic. Crops moved
from their center of origin to a wide range of environments, where natural
selection drove genetic adaptation to the new ones. Equally important are the genetic
structures associated with end-use or cultural preferences that lead to the
increase of the frequency of favorable alleles. Although they might become fixed
within populations, would still be polymorphic in worldwide collections of
cultivars or landraces and should be characterized as QTL in mapping studies of
diverse material (Hamblin et al., 2011).
PRACTICAL APPLICATIONS OF DEEP GENOTYPING
As discussed, the application of
deep genotyping data in the genetic characterization of a germplasm base is
important to assess genetic diversity, genetic relatedness, and population
structure, contributing to a better understanding of the materials included in
a breeding program. One molecular breeding application requiring high-density
markers is genomic selection (GS). Although it is not currently used routinely,
its importance and consideration are clearly growing. The great advantage of GS
use is the ability to accurately select individuals of higher breeding value
without the requirement of collecting phenotypes pertaining to these
individuals. This can facilitate a shortening of the breeding cycle and enable
rapid selection and intercrossing of early-generation breeding material.
GS consists of the prediction of
the genomic estimated breeding value (GEBV) of individuals based on genomic data
(Meuwissen et al., 2001). Typically, is performed among the progeny of a
biparental cross between two elite inbreds (breeding population) where
phenotypes and genome-wide genotypes are investigated in the training
population (a subset of the breeding population) to predict significant
relationships between phenotypes and genotypes using statistical approaches.
Marker effects estimated on the training population will be used to predict the
performance of the best candidates in the rest of the breeding population
solely based on GEBV (Daetwyler et al., 2013; Heslot et al.,
2015). One question that arises is: how many SNP loci should be genotyped to
achieve a reasonable prediction accuracy (e.g., 0.6 correlation between true
breeding value and GEBV)? There is no single answer, however there are some
aspects to consider that can bring us closer to it. Simulation studies showed
that the relationships between the individuals in the training population and the
individuals in the prediction population had a major impact on the accuracy of
the GEBV. Accurate predictions could be obtained with a small number of markers
(e.g., 300–500) and a small number of phenotypes (e.g., 200– 1000) when the
phenotypes were collected from closely related biparental populations. To
generate accurate predictions from nominally unrelated individuals many more
phenotypes (e.g., 20,000) and many more markers (e.g., 10,000) were required
(Hickey et al., 2014). GS provides tremendous opportunities to increase
genetic gain in plant breeding. Early empirical and simulation results are
promising, but for GS to work, consideration of the cost-benefit balance is
needed.
Although deep genotyping allows
for the identification of thousands of informative SNPs, most routine molecular
breeding applications do not require such a large number of markers. Therefore,
the selection of a subset of SNP markers suitable for the chosen breeding strategy
and their conversion to a more cost-effective genotyping technology is
recommended. Kompetitive Allele Specific PCR (KASP) is a user-friendly SNP
platform that is cost efficient for smaller numbers of markers (<200) which
is what is needed for marker-assisted recurrent selection, marker-assisted
backcrossing, and quality control analysis. KASP is one of the uniplex SNP genotyping
platforms that has evolved to be a global benchmark technology for conversion
of selected SNP (Semagn et al., 2014). Practical applications that
require around 200 markers include quality control analysis (genetic identity,
genetic purity, and parentage verification), linkage mapping of QTL,
marker-assisted recurrent selection, and markerassisted backcrossing.
Quality control
Control of the genetic purity (in
terms of identity of the parental inbred lines and progeny testing of the resulting
F1 hybrids) is an essential quality control (QC) parameter in hybrid breeding,
as maintaining high levels of genetic purity is critical to guarantee a robust
and stable agronomic performance of the genotype. Genetic purity evaluation is
also relevant to meeting the strict intellectual property requirements that
govern plant breeding and variety registration in many countries (Chen et
al., 2016; Josia et al., 2021). Genetic purity can be proved using
different approaches such as grow out test, use of biochemical markers and use
of molecular markers. The grow out test is based on the use of a set of
morphological descriptors and the biochemical marker approach analyzes
electrophoretic protein (isoenzymes) profiles. Molecular marker approaches detect
the variation of genotypes directly at the DNA level and have several
advantages including high polymorphism, high-throughput detection methods, and
they are unaffected by environmental conditions or the physiological stage of
the plant (Chen et al., 2016; Josia et al., 2021). The main
purpose of routine QC genotyping is to identify contamination or mislabeling of
germplasm during regeneration, seed increase or seed distribution. To achieve a
cost-effective QC test, a balance between accuracy of detection and efficiency needs
to be maintained, for which optimization of the balance between accuracy and
cost is the main concern when choosing a set of markers for QC.
In maize, was proposed the use of
two separate sets of markers, each focusing on different types of QC. The first
was a broad QC focusing on identity of a sample employing a minimum of 80 KASP
markers (which were selected based on MAF, coverage and chromosome distribution)
to distinguish each of the entries from one another. It is important to conduct
this type of QC before starting new breeding crosses to ensure the identity and
purity of the founding parents and to evaluate the levels of residual
heterogeneity within them. The second approach was rapid QC for seed production
using a smaller sub-set of only ten selected KASP markers (Chen et al.,
2016).
QTL mapping and marker-assisted recurrent selection
The nature of a trait may sometimes
suggest that much of the quantitative variation is controlled by a few genes with
large effects. In this situation, the objective of QTL mapping is finding a few
major QTL. The subsequent breeding strategy is to introduce or pyramid these
QTL, via standard breeding procedures, into elite germplasm to develop improved
cultivars. Exploiting a few major QTL therefore requires both gene discovery
(i.e., QTL mapping) and selection (Bernardo, 2008).
QTL mapping involves
identification of a subset of markers that are significantly associated with
one or more QTL influencing the expression of the trait of interest. The main
steps in linkage-based QTL mapping include (1) selecting and/or developing
appropriate biparental mapping populations; (2) phenotyping the population for
the trait of interest under greenhouse and/or field conditions; (3) choosing
the molecular marker system, genotyping the parents of the mapping population
and F1 with larger numbers of markers, and selecting markers exhibiting
polymorphism between the parents; (4) choosing a genotyping approach (entire population,
selective genotyping, or bulk segregant analysis) and generating molecular data
for an adequate number of uniformly-spaced polymorphic markers; and (5)
identifying the molecular markers associated with the QTL using statistical
programs (Semagn et al., 2010, 2014). There is no clear consensus
regarding the number of markers demanded for genotyping bi-parental populations
but depending on the species and its genetic map, most researchers use around
200 and 400 markers. Once a significant QTL is identified, a second round of
genotyping can be performed by saturating the chromosome region with additional
polymorphic SNPs around the QTL of interest (fine mapping). Chromosome position
of the QTL will be established relative to closely spaced flanking SNPs, and
these markers can potentially be used for marker assisted selection (MAS) of
the QTL associated to the trait.
The nature of a trait may
sometimes suggest that much of the quantitative variation is controlled by many
genes with small effects. Two related approaches have been proposed and used to
increase the frequency of favorable QTL alleles at multiple loci: (i) F2
enrichment followed by inbreeding and (ii) marker-assisted recurrent selection
(MARS) (Bernardo, 2008). MARS refers to the improvement of an F2 population by
one generation of phenotypic selection in the target set of environments
followed by 2–3 generations of selection based on significant marker genotypes.
MARS has been applied for improving a breeding population with respect to QTLs
exerting smaller effects on the phenotype (Gokidi et al., 2016).
In both approaches the base
generation is usually an F2 population from the cross between two inbreds, although
backcrosses, three-way crosses, or double crosses may also be used. The
objective is to develop a recombinant inbred with superior per se performance
for self-pollinated crops or with superior testcross performance for hybrid
crops. Whereas F2 enrichment usually involves only one generation of
marker-based selection, MARS involves several cycles of marker-based selection
(Bernardo, 2008).
Marker-assisted backcrossing
Marker-assisted backcrossing
(MABC) is used for transferring genes which are responsible for favorable agronomic
traits from a donor line into the genome of a recipient (recurrent) line.
Introgression of a QTL by successive backcrosses is used to improve elite lines
(recurrent parent) by introducing alleles from exotic material (donor parent).
Besides to maintain the donor allele at the QTL in the progenies, the process
pursues two objectives: reduction of the size of the donor genetic background
around the target locus, and recovery of the recurrent parent genetic
background (Hospital, 2005).
In the absence of selection, the
proportion of the donor genome decreases by half at each generation. Thus, it is
expected that after five backcross generations (BC5), 98.4% of recurrent parent
background is recovered. However, since selection is for the donor allele at
the QTL, elimination of the donor genome around that QTL will be much slower
than in the rest of the genome. As a result, the proportion of the donor genome
will decrease less for the chromosome carrying the target locus than for the
others. This is the so-called linkage drag problem (Naveira and Barbadilla,
1992).
Marker-assisted selection (MAS)
in introgression of favorable alleles at QTL usually comprises selection for presence
of the donor allele at two markers delimiting the interval in which the
putative QTL was detected, and the recurrent parent allele at markers outside
the QTL interval (foreground selection). The use of tightly-linked flanking
markers for recurrent parental alleles helps to decrease linkage drag more
rapidly resulting in short donor chromosome segments attached to the target gene.
To optimize the positions of a limited number of markers that flank the target
locus was concluded that the larger the population, the closer the markers
should be to the target locus (Frisch and Melchinger, 2005).
Marker distance and distribution
for genome-wide background selection will impact significantly on the efficiency
of MABC method. Contrary to common belief, high marker densities are not
required. To efficiently identify the backcross individuals with the smallest percentage
of donor genome, a marker distance of 10 cM is sufficient. Decreasing the
marker distances below 10 cM had only marginal effect on the recipient genome recovery.
One explanation for this result is that, in general, one crossing over by
meiosis and chromatid occurs for each chromosome segment 1 M in length. In two-
or three-generation backcrossing programs, the number of recombination events
resulting in chromosome segments of different parental origin is therefore
limited (Herzog and Frisch, 2011, 2013). Computer simulations were conducted to
evaluate and optimize the resource requirements of conversion programs of
different crop genetic models with chromosome numbers (from n=7 to n=17)
demonstrating how MABC contributes to reduce the time and costs demanded for
gene introgression. The results showed that depending on the genome size of the
crop of interest, recovering 10% quantile with 98% of recurrent background can
be reach in BC3 working with population sizes comprised between 10 to 30
individuals per generation and around two to three SNP markers per chromosome
equally distributed across each linkage group (Herzog and Frisch, 2013). A
further considerable reduction of the costs could be achieved if the population
size in the first backcross generation is twice the population size in
generations BC2 and BC3 of a three-generation backcrossing program (Herzog and Frisch,
2013).
CONCLUSIONS
One of the greatest challenges
facing humanity is the development of sustainable strategies to ensure food availability
in response to population growth and climate change. Different foresight
studies have concurrently argued that current food production practices would not
be sufficient and therefore a transformation of the food system is required.
One approach that can contribute to increase food security is to close yield
gaps and enhancing genetic gain, for which solutions based on multiple
disciplines should be found. Among these, clearly the genetic improvement of
crops and specifically molecular breeding plays a fundamental role.
There is no doubt that a crop
breeding program is fundamentally based on the quality of the germplasm. However,
if a detailed genetic characterization is not available, there is a risk of
underusing genetic resources or delaying the development of superior varieties.
As stated, deep genotyping is an essential tool for a comprehensive
characterization of the germplasm of interest and, fortunately, the technology
is now accessible at a reasonable cost. What must be ensured is the correct
interpretation of the genotypic information and on that basis develop efficient
crop breeding strategies that respond to the real needs of the breeding program.
REFERENCES
Bailey-Serres J., Parker J.E.,
Ainsworth E.A., Oldroyd G.E.D., Schroeder J.I. (2019) Genetic strategies for
improving crop yields. Nature 575: 109-118.
Beckett T.J., Morales A.J.,
Koehler K.L., Rocheford T.R. (2017) Genetic relatedness of previously
Plant-Variety-Protected commercial maize inbreds. PLoS One 12: e0189277.
Bernardo R. (2008) Molecular
markers and selection for complex traits in plants: Learning from the last 20
years. Crop Sci. 48: 1649-1664.
Bohar R., Chitkineni A., Varshney
R.K. (2020) Genetic molecular markers to accelerate genetic gains in crops.
Biotechniques 69: 158-160.
Buckler E.S., Thornsberry J.F.,
Kresovich S. (2001) Molecular diversity, structure and domestication of
grasses. Genet.
Res. 77: 213–218.
Chen
J, Zavala C, Ortega N, Petroli C, Franco J, Burgueño J, Costich DE; Hearne SJ. (2016) The development of quality control genotyping approaches: a case
study using elite maize lines. PLoS ONE 11(6): e0157236.
https://doi.org/10.1371/journal.pone.0157236.
Clark R.M., Linton E., Messing
J., Doebley J.F. (2004) Pattern of diversity in the genomic region near the
maize domestication gene tb1. Proc. Natl. Acad. Sci. USA 101: 700-707.
Cobb J.N., Juma R.U., Biswas
P.S., Arbelaez J.D., Rutkoski J., Atlin G., Hagen T., Quinn M., Ng E.H. (2019)
Enhancing the rate of genetic gain in public-sector plant breeding programs: lessons
from the breeder’s equation. Theor. Appl. Genet. 132: 627-645.
Cooper M., Messina C.D., Podlich
D., Totir L.R., Baumgartern A., Hausmann N.J., Wright D., Graham G. (2014)
Predicting the future of plant breeding: complementing empirical evaluation
with genetic prediction. Crop Pasture Sci. 65: 311–336.
Darrier B., Russell J., Milner
S.G., Hedley P.E., Shaw P.D., Macaulay M., Ramsay L.D., Halpin C., Mascher M.,
Fleury D.L., Langridge P., Stein N., Waugh R.A. (2019) Comparison of mainstream
genotyping platforms for the evaluation and use of barley genetic resources.
Front. Plant Sci. 10: 544. doi: 10.3389/fpls.2019.00544.
Daetwyler H.D., Calus M.P.L.,
Pong-Wong R., de los Campos G., Hickey J.M. (2013) Genomic prediction in
animals and plants: simulation of data, validation, reporting, and
benchmarking. Genetics 193: 347–365.
Dempewolf H., Baute G., Anderson
J., Kilian B., Smith C., Guarino L. (2017) Past and future use of wild
relatives in crop breeding. Crop Sci. 57: 1070-1082.
Ellis D.; Chavez O.; Coombs J.;
Soto J.; Gomez R.; Douches D.; Panta A.; Silvestre R.; Anglin N.L. (2018)
Genetic identity in genebanks: Application of the SolCAP 12K SNP array in fingerprinting
and diversity analysis in the global in trust potato collection. Genome 61: 523–537.
Elshire R.J., Glaubitz J.C., Sun
Q., Poland J.A., Kawamoto K., Buckler E.S., Mitchell S.E. (2011) A robust,
simple genotypingby- sequencing (GBS) approach for high diversity species. PLoS
One 6: e19379.
Ertiro B.T., Semagn K., Das B.,
Olsen M., Labuschagne M., Worku M., Wegary D., Azmach G., Ogugo V., Keno T.,
Abebe B., Chibsa T., Menkir A. (2017) Genetic variation and population
structure of maize inbred lines adapted to the mid-altitude sub-humid maize
agro-ecology of Ethiopia using single nucleotide polymorphic (SNP) markers. BMC
Genomics 18: 777. https://doi.org/10.1186/ s12864-017-4173-9.
FAO (1997) The state of the
world’s plant genetic resources for food and agriculture. Food and Agriculture
Organization of the United Nations, Rome, Italy.
Flint-Garcia S.A., Thornsberry
J.M, Bukler, E.S. (2003) Structure of linkage disequilibrium in plants. Annu.
Rev. Plant Biol. 54: 357-374.
Flint-Garcia S.A., Buckler E.S.,
Tiffin P., Ersoz E., Springer N.M. (2009) Heterosis is prevalent for multiple
traits in diverse maize germplasm. PLoS One 4: e7433.
Frascaroli E., Schrag T.A.,
Melchinger A.E. (2013) Genetic diversity analysis of elite European maize (Zea
mays L.) inbred lines using AFLP, SSR, and SNP markers reveals ascertainment
bias for a subset of SNPs. Theor Appl Genet. 126: 133–141.
Frisch M., Melchinger A.E. (2005)
Selection theory for marker-assisted backcrossing. Genetics 170: 909-917.
Fu Y.-B., Cober E.R., Morrison
M.J., Marsolais F., Peterson G.W., Horbach C. (2021) Patterns of genetic
variation in a soybean germplasm collection as characterized with genotypingby-
sequencing. Plants 10(8): 1611. https://doi.org/10.3390/plants100816111611.
Garris A.J., McCouch S.R.,
Kresovich S. (2003) Population structure and its effects on haplotype diversity
and linkage disequilibrium surrounding the xa5 locus of rice (Oryza
sativa L.). Genetics 165: 759–769. Goddard M. (2009) Genomic selection: prediction
of accuracy and maximisation of long term response. Genetica 136: 245-257.
Gokidi, Y., Bhanu, A. N., Singh,
M. N. (2016) Marker assisted recurrent selection: an overview. Adv. Life Sci.
5: 6493–6499.
Govindaraj M., Vetriventhan M.,
Srinivasan M. (2015) Importance of genetic diversity assessment in crop plants
and its recent advances: an overview of its analytical perspectives. Genet.
Res. Int. 2015: 431487. doi: 10.1155/2015/431487.
Hamblin M.T., Buckler E.S.,
Jannink J.L. (2011) Population genetics of genomics-based crop improvement
methods. Trends Genet. 27: 98-106.
Heslot N., Jannink J.L., Sorrells
M.E. (2015) Perspectives for genomic selection applications and research in
plants. Crop Sci. 55: 1–12.
Herzog E., Frisch, M. (2011)
Selection strategies for marker-assisted backcrossing with high-throughput
marker systems. Theor. Appl. Genet. 123: 251–260.
Herzog E., Frisch M. (2013)
Efficient markerassisted backcross conversion of seedparent lines to
cytoplasmic male sterility. Plant Breeding 132: 33–41.
Hospital F. (2005) Selection in
backcross programs. Philos. Trans. R Soc. Lond. B Biol. Sci. 360: 1503–11.
Hickey J.M., Dreisigacker S.,
Crossa J., Hearne S., Babu R., Prasanna B.M., Grondona M., Zambelli A.,
Windhausen V.S., Mathews K., Gorjanc G. (2014) Evaluation of genomic selection
training population designs and genotyping strategies in plant breeding programs
using simulation. Crop Sci. 54: 1476–1488.
Hickey L.T., Hafeez Amber N.,
Robinson H., Jackson S.A., Leal-Bertioli S.C.M., Tester M., Gao C., Godwin
I.D., Hayes B.J., Wulff B.B.H. (2019) Breeding crops to feed 10 billion. Nat. Biotechnol.
37: 744-754.
Ismail A.M., Horie T. (2017)
Genomics, physiology, and molecular breeding approaches for improving salt
tolerance. Annu. Rev. Plant Biol. 68: 405-434.
Josia C., Mashingaidze K.,
Amelework A.B., Kondwakwenda A., Musvosvi C., Sibiya J. (2021) SNP-based
assessment of genetic purity and diversity in maize hybrid breeding. PLoS ONE
16(8): e0249505. https://doi. org/10.1371/journal.pone.0249505.
Jeong N., Kim K.-S., Jeong S.,
Kim J.-Y., Park S.-K., Lee J.S., Jeong S.-C., Kang S.-T., Ha B.-K., Kim D.-Y.,
Kim N., Moon J.-K, Choi M.S. (2019) Korean soybean core collection: Genotypic
and phenotypic diversity population structure and genome-wide association
study. PLoS One 14: e0224074.
Labroo M.R., Studer A.J.,
Rutkoski J.E. (2021) Heterosis and hybrid crop breeding: a multidisciplinary
review. Front. Genet. 12: 643761. doi:10.3389/fgene.2021.643761.
Lachance J., Tishkoff S.A. (2013)
SNP ascertainment bias in population genetic analyses: why it is important, and
how to correct it. BioEssays 35: 780–786.
Lam H.M., Xu X., Liu X., Chen W.,
Yang G., Wong F.L., Li M.W., He W., Qin N., Wang B., Li J., Jian M., Wang J.,
Shao G., Wang J., Sun S.S., Zhang G. (2010) Resequencing of 31 wild and
cultivated soybean genomes identifies patterns of genetic diversity and
selection. Nat. Genet. 42: 1053–1059.
Leitão S.T., Dinis M., Veloso
M.M., Šatović Z., Vaz Patto M.C. (2017) Establishing the bases for
introducing the unexplored portuguese common bean germplasm into the breeding world.
Front. Plant Sci. 8: 1296. doi: 10.3389/fpls.2017.01296.
Lenaerts B., Collard B.C.Y.,
Demont M. (2019) Improving global food security through accelerated plant
breeding. Plant Sci. 287: 110207.
Li H., Vikram P., Singh R.P.,
Kilian A., Carling J., Song J., Burgueno-Ferreira J.A., Bhavani S.,
Huerta-Espino J., Payne T., Sehgal D., Wenzl P., Singh S. (2015) A high density
GBS map of bread wheat and its application for dissecting complex disease
resistance traits. BMC Genomics 16: 216.
https://doi.org/10.1186/s12864-015-1424-5.
Liu A., Burke J.M. (2006)
Patterns of nucleotide diversity in wild and cultivated sunflower. Genetics
173: 321-330.
Lucek K., Willi Y. (2021) Drivers
of linkage disequilibrium across a species’ geographic range. PLoS Genet. 17:
e1009477.
Meuwissen T.H.E., Hayes B.J.,
Goddard M.E. (2001) Prediction of total genetic value using genome-wide dense
marker maps. Genetics 157: 1819–1829.
Milner S.G., Jost M., Taketa S.,
Mazon E R., Himmelbach A., Oppermann M., Weise S., Knüpffer H., Basterrechea
M., König P., Schüler D., Sharma R., Pasam R.K., Rutten T., Guo G., Xu D.,
Zhang J., Herren G., Müller T., Krattinger S.G., Keller B., Jiang Y., González M.Y.,
Zhao Y., Habekuß A., Färber S., Ordon F., Lange M., Börner A., Graner A., Reif
J.C., Scholz U., Mascher M., Stein N. (2019). Genebank genomics reveals the
diversity of a global barley collection. Nat. Genet. 51: 319–326.
Mohammadi S.A., Prasanna B.M.
(2003) Analysis of genetic diversity in crop plants - salient statistical tools
and considerations. Crop Sci. 43: 1235–1248.
Mondini L., Noorani A., Pagnotta
M.A. (2009) Assessing plant genetic diversity by molecular tools. Diversity 1:
19-35.
Naveira H., Barbadilla A. (1992)
The theoretical distribution of lengths of intact chromosome segments around a
locus held heterozygous with backcrossing in a diploid species. Genetics 1992:
130:205–209.
Negro S.S., Millet E.J., Madur
D., Bauland C., Combes V., Welcker C., Tardieu F., Charcosset A., Nicolas S.D.
(2019) Genotypingby- sequencing and SNP-arrays are complementary for detecting
quantitative trait loci by tagging different haplotypes in association studies.
BMC Plant Biol. 19: 318. https://doi.org/10.1186/s12870-019-1926-4.
Nordborg M. (2000) Linkage
disequilibrium, gene trees and selfing: an ancestral recombination graph with
partial selffertilization. Genetics 154: 923–929.
Pavan S., Delvento C., Ricciardi
L., Lotti C., Ciani E., D’Agostino N. (2020) Recommendations for choosing the
genotyping method and best practices for quality control in crop genomewide association
studies. Front. Genet. 11: 447. doi: 10.3389/fgene.2020.00447.
Pourkheirandish M., Golicz A.A., Bhalla
P.L., Singh M.B. (2020) Global role of crop genomics in the face of climate
change. Front. Plant Sci. 11: 922. doi: 10.3389/fpls.2020.00922. Pritchard
J.K., Stephens M., Rosenberg N.A., Donnelly P. (2000) Association mapping in structured
populations. Am. J. Hum. Genet. 67: 170–181.
Rasheed A., Hao Y., Xia X., Khan
A., Xu Y., Varshney R. K., He Z. (2017) Crop breeding chips and genotyping
platforms: progress, challenges, and perspectives. Mol. Plant. 10: 1047–1064.
Reif J.C., Melchinger A.E., Xia X.C.,
Warburton M.L., Hoisington D.A., Vasal S.K., Beck D., Bohn M., Frisch M. (2003)
Use of SSRs for establishing heterotic groups in subtropical maize. Theor.
Appl. Genet. 107: 947–957.
Remington D.L., Thornsberry J.M.,
Matsuoka Y., Wilson L.M., Whitt S.R., Doebley J., Kresovich S., Goodman M.M.,
Buckler E.S. (2001) Structure of linkage disequilibrium and phenotypic
associations in the maize genome. Proc. Natl. Acad. Sci. USA 98: 11479–11484.
Sansaloni C., Franco J., Santos
B., Percival- Alwyn L., Singh S., Petroli C., Campos J., Dreher K., Payne T.,
Marshall D., Kilian B., Milne I., Raubach S., Shaw P., Stephen G., Carling J.,
Pierre C.S., Burgueño J., Crosa J., Li H., Guzman C., Kehel Z., Amri A., Kilian
A., Wenzl P., Uauy C., Banziger M., Caccamo M., Pixley K. (2020) Diversity
analysis of 80,000 wheat accessions reveals consequences and opportunities of
selection footprints. Nat. Commun. 11: 4572.
https://doi.org/10.1038/s41467-020-18404-w
Scheben A., Batley J., Edwards D.
(2017) Genotyping-by-sequencing approaches to characterize crop genomes:
choosing the right tool for the right application. Plant Biotechnol. J.
15:149-161.
Semagn, K, Bjørnstad, Å, Xu, Y.
(2010) The genetic dissection of quantitative traits in crops. Electron. J.
Biotechnol. 13: 16-17.
Semagn, K., Babu, R., Hearne, S.:
Olsen M (2014) Single nucleotide polymorphism genotyping using Kompetitive
Allele Specific PCR (KASP): overview of the technology and its application in
crop improvement. Mol. Breeding 33: 1–14.
Silva K.J., Pastina M.M.,
Guimarães C.T., Magalhães J.V., Pimentel L.D., Schaffert R.E., Pinto M.O.,
Souza V.F., Bernardino K.C., Silva M.J., Borém A., Menezes C.B. (2021) Genetic diversity
and heterotic grouping of sorghum lines using SNP markers. Sci. Agric. 78: e20200039.
Singh N., Wu S., Raupp W.J.,
Sehgal S., Arora S., Tiwari V., Vikram P., Singh S., Chhuneja P., Gill B.S.,
Poland J. (2019) Efficient curation of genebanks using next generation sequencing
reveals substantial duplication of germplasm accessions. Sci. Rep. 9: 650. doi:
10.1038/s41598-018-37269-0.
Slatkin M. (2008) Linkage
disequilibrium— understanding the evolutionary past and mapping the medical
future. Nat. Rev. Genet. 9: 477–485.
Smith S., Bubeck D., Nelson B.,
Stanek J., Gerke J. (2015) Genetic diversity and modern plant breeding. In:
Ahuja M., Jain S. (Eds.) Genetic diversity and erosion in plants. Sustainable Development
and Biodiversity. Springer, Cham, pp. 55-88.
Varshney R.K., Graner A.,
Sorrells M.E. (2005) Genomics-assisted breeding for crop improvement. Trends
Plant Sci. 10: 621–630.
Varshney R.K., Bohra A., Yu J.,
Graner A., Zhang Q., Sorrells M.E. (2021) Designing future crops:
genomics-assisted breeding comes of age. Trends Plant Sci. 26: 631-649.
Vendelbo N.M., Sarup P., Orabi
J., Kristensen P.S., Jahoor A. (2020) Genetic structure of a germplasm for
hybrid breeding in rye (Secale cereale L.). PLoS One 15: e0239541.
Wang J. (2016) Pedigrees or
markers: Which are better in estimating relatedness and inbreeding coefficient?
Theor. Popul. Biol. 107: 4-13
Whitt S.R., Wilson L.M.,
Tenaillon M.I., Gaut B.S., Buckler E.S. (2002) Genetic diversity and selection
in the maize starch pathway. Proc. Natl. Acad. Sci. USA 99: 12959-12962.
Wu X., Li Y., Shi Y., Song Y.,
Wang T., Huang Y., Li Y. (2014) Fine genetic characterization of elite maize
germplasm using highthroughput SNP genotyping. Theor. Appl. Genet. 127:
621-631.
Xu S., Zhu D., Zhang Q. (2014)
Predicting hybrid performance in rice using genomic best linear unbiased
prediction. Proc. Natl. Acad. Sci. USA 111: 12456–12461.
Xu Y., Li P., Zou C., Lu Y., Xie
C., Zhang X., Prasanna B.M., Olsen M.S. (2017) Enhancing genetic gain in the
era of molecular breeding. J. Exp. Bot. 68: 2641-2666.
Zambelli A. (2019) The impact of
molecular genetics in plant breeding: realities and perspectives. J. Basic
Appl. Genet. 30: 11-15.
Zhao Y., Li Z., Liu G., Jiang Y.,
Maurer H.P., Wurschum T., Mock H.P., Matros A., Ebmeyer E., Schachschneider R.,
Kazman E., Schacht J., Gowda M., Longin C.F., Reif J.C. (2015) Genome-based
establishment of a high-yielding heterotic pattern for hybrid wheat breeding.
Proc. Natl. Acad. Sci. USA 112: 15624–15629.
Zhu C., Gore M., Buckler E.S., Yu
J. (2008) Status and prospects of association mapping in plants. Plant Genome
1: 5-20.
Zhu Y.L., Song Q.J., Hyten D.L.,
Van Tassell C.P., Matukumalli L.K., Grimm D.R., Hyatt S.M., Fickus E.W., Young
N.D., Cregan P.B. (2003) Single-nucleotide polymorphisms in soybean. Genetics
163: 1123–1134.



